Utah NeoSeq Project
University of Utah Health launched a pilot project with a goal of quickly and accurately detecting genetic disorders among newborns in its neonatal intensive care unit (NICU). Clinicians believe that bringing precision medicine to the NICU will transform neonatal care, leading to better diagnosis and treatment of critically ill infants with genetic-related diseases while lowering medical costs and easing the emotional and financial burden on parents and families. With this project, U of U Hospital will join Primary Children’s Hospital, also in Salt Lake City, as being part of a small group of hospitals worldwide to offer this type of advanced care.

Electronic Health Record (EHR)
Dr. Yandell is combining clinical variables with sequence data for discovery and outcomes research. As part of this project, he seeks to create a best in class workflow for screening NICU patients to identify best candidates for rapid whole genome sequencing. This project also provides the general means for outcomes research that can combine sequence data with clinical variables, e.g. diagnoses, procedures, medications and labs. The goals of Dr. Yandel’s EHR project are to create infrastructure and tools for assembly of maximally informative cohorts using Utah’s unique data resources, and enable Dr. Yandell to continue to develop the CAE and WARP tools for mining the Utah Population Database and Electronic Data Warehouse for the purposes of precision medicine.

Genomic Data Sharing Portal
Dr. Gabor Marth’s team and the UCGD Core have been developing a genomic data sharing portal for the University of Utah. This web-based resource allows collaborative access, sharing, and exploration of genomic data among both internal and eventually external investigators. The portal enables the organization and management of genomic data projects, provides administrative support/dashboards for projects, and supports clinician driven genomic analysis. The goals of this project are to enable Dr. Marth and the UCGD Core to continue the development and implementation of the portal.

Precision Oncology Project
Drs. Gabor Marth and Aaron Quinlan from UCGD have partnered with Drs. Bryan Welm and Alana Welm to conduct a precision oncology pilot project. This project seeks to integrate functional drug screens with multi-omic molecular profiling of the patient tumor to provide the physician with clinically actionable lines of evidence – which drugs worked and why they worked – during the patient’s treatment timeline. The project will establish cutting-edge approaches to the care of advanced cancers and establish a foundation of infrastructure, technology, and operating procedures for precision oncology at the University of Utah.

Long-Read Sequencing
Dr. Aaron Quinlan has been developing an innovative method to efficiently enrich for and sequence disease genes in structurally complex genomic regions via nanopore and PacBio sequencing instruments. In the context of rare disease diagnosis, ~50% of patients go undiagnosed, in large part owing to an inherent limit to variant discovery with Illumina sequencing. Very long, accurate DNA reads from the PacBio HiFi technology overcome this limit, and is especially effective for detecting (1) tandem repeat expansions that are larger than the Illumina read lengths; (2) for structural variants (SVs) whose breakpoints often lie in repeat-rich sequence; and (3) for using the long reads to phase distant variants and better detect compound heterozygotes. Notably, more than half of the existing structural variants and repeat expansions >50bp are missed by Illumina whole genome sequencing (WGS), yet recovered by PacBio. His team work with ARUP Laboratories to evaluate the potential of PacBio HiFi sequencing’s potential for Mendelian disorder discovery.

Multi-Omics and Metagenomics Integration for Genetic Diagnosis
The UCGD investigators established dedicated teams for innovation and integration of multi-omics approaches into current workflows and projects for better discovery power in all disease focus areas. The goal of this project is to develop algorithms for the utilization of long-read sequencing data, which will increase discovery power for NeoSeq, Penelope/UDN, and support the UCGD’s new center grant proposals (e.g., Mendelian Genomics Research Centers and Centers of Excellence in Genomic Science).

New algorithms and tools for large-scale genomic analyses
The exploration and interpretation of large, complex datasets is vital to discovery in genomics. However, researchers now confront a fundamental limitation; unprecedented experiments are possible thanks to modern DNA sequencing technologies, yet existing “genome arithmetic” algorithms and data formats for comparing and dissecting the resulting datasets are incapable of keeping pace with inexorable growth in dataset size and complexity. The overall objective of this project is to provide the genomics community with innovative new algorithms and software that keep pace with modern genomics experiments and facilitate future discoveries.

Calypso: a web software system supporting team-based, longitudinal genomic diagnostic care
Some of the most challenging diagnostic cases involve patients who present with complex phenotypes that evolve over time and intersect multiple medical disciplines. We are developing software to support team-based, long-term genomic analysis in challenging settings such as undiagnosed disease clinics and rapid genome sequencing programs to rapidly diagnose critically ill newborns in neonatal intensive care units called Calypso. UCGD team put Calypso into the hands of the clinical teams in our undiagnosed disease clinic and our neonatal intensive care unit, ideal testing grounds for evaluating our tool’s impact on long-term genome diagnostic care in the context of team-based medicine. We are planning to deploy and evaluate our tool in NHGRI’s Undiagnosed Disease Network to help realize the collaborative diagnostic opportunities presented by a large, clinically focused research consortium.

Scalable detection and interpretation of structural variation in human genomes
Structural variation (SV), is a diverse class of genome variation that includes copy number variants (CNVs) such as deletions and duplications, as well as balanced rearrangements, such as inversions and reciprocal translocations. A typical human genome harbors >4,000 SVs larger than 300bp and their large size increases the potential to delete or duplicate genes, disrupt chromatin structure, and alter expression. Despite their prevalence and potential for phenotypic consequence, SVs remain notoriously difficult to detect and genotype with high accuracy. Much of this difficulty is driven by the fact DNA sequence alignment “signals” indicating SVs are far more complex than for single-nucleotide and insertion deletion variants.
SEMIColon: Somatic Exploration of Mosaicism In Colon
Somatic mosaicism refers to the accumulation of mutations in non-germline cells throughout an organism’s lifetime. Somatic mutations exist in healthy tissues and play a central role in disease pathogenesis, from developmental syndromes to cancer. In the colon, somatic mosaicism is a causative factor in colorectal cancer (CRC) and has been proposed to contribute to inflammatory bowel disease (IBD). The Center for Genomic Medicine endeavors to identify better diagnoses, treatments, and prevention strategies. This project aims to identify regional baselines of healthy colonic mutational landscapes, a key step needed to establish guidelines for improved screening and surveillance strategies for colorectal pathologies early on in pathogenesis, when intervention or disease mitigation is possible. This advance would reduce the morbidity and mortality of colorectal pathologies by limiting their progression prior to recognition and treatment.
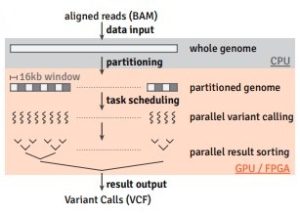
Accelerating genomic analysis for time critical clinical applications
Lead Investigators: Dr. Gabor Marth and Dr. Yi Qiao
The current next generation sequencing (NGS) data analysis turnaround, not sequencing itself, is too slow for time sensitive clinical applications, such as identifying the optimal treatment for a cancer patient days after biopsying, and according to the unique genomic characteristics of that patient’s tumor. This exploratory development proposal aims to greatly accelerate genomic analysis (60X whole human genome in 10 minutes) using heterogeneous computing techniques. In addition to enabling NGS for time critical clinical applications, our emphasis on an open-source development model, reusable software components, and the acceleration of widely used algorithms will allow for the adoption of accelerated computing by a large developer and user community.
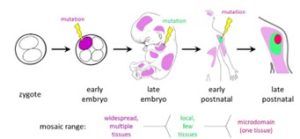
A reference-free computational algorithm for comprehensive somatic mosaic mutation detection.
Lead Investigators: Dr. Gabor Marth and Dr. Hunter Underhill
Somatic mosaicism (SM), i.e. the presence of cells with somatically acquired mutations, is a driving feature of cancer and several developmental diseases. However, whereas today we have detailed understanding and predictive models of benign and pathogenic inherited polymorphisms, germline de novo mutations, and tumor mutations, we have only limited knowledge of the burden, allele frequency behavior, clonal patterns, and mutational signatures of healthy somatic mosaicism. Realizing that such currently missing knowledge is critical for informing experimental design in future studies of mosaicism’s biological and clinical consequences, NIH is launching an ambitious initiative, the Somatic Mosaicism across Human Tissues (SMaHT) project to construct a comprehensive human somatic mosaicism atlas.
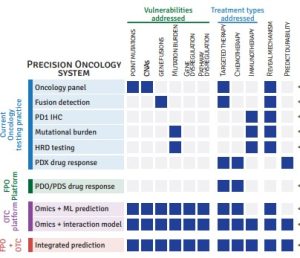
Functional precision oncology to understand and predict treatment response in solid tumors.
Lead Investigators: Gabor Marth D.Sc. (contact), Alana Welm, PhD, Sam Cheshier MD/PhD
The overall goal of this project is to pilot whole-genome CRISPR dropout screens as a powerful approach for comprehensively identifying critical gene vulnerabilities of individual patient tumors. The resulting list of critical vulnerabilities will then be used to validate computational predictions (in PPG Project 1); guide therapeutic targeting in PPG Projects 2 & 3); and study the evolution of tumor vulnerabilities over time (in refractory tumors, PPG Project 3) and space (across metastatic sites PPG Project 2).
