Genomic Medicine

Genetics of Inherited Disease
We analyze genome sequences to determine the genetic cause of a broad range of inherited disorders, including heart and lung disease, familial cancers, inflammatory and immune conditions, metabolic disease, and neurological conditions.
We participate in large-scale genome sequencing projects including the Utah Genome Project, the Pediatric Cardiac Genomics Consortium, the Simon’s Foundation Autism Research Initiative, and the Sudden Cardiac Death in the Young Consortium. We also partner with life sciences and technology companies to develop novel software tools, diagnostics, and precision therapies based on our research.
Browse our inherited disease publications here.
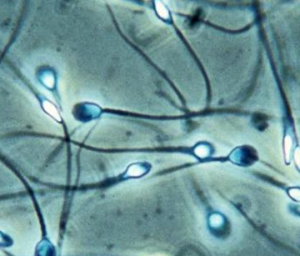
What are the connections between sperm mutation, rates and disease, subfertility and aging?
Mutations in sperm and egg drive human evolution, yet also underlie human disease. Their rates, patterns, and underlying mechanisms are therefore essential to understand. Work from our group and others have genome sequenced blood from parent-child trios to find that sperm contribute 80% of transmissible mutations. However, these pedigree analyses study mutations solely from sperm capable of facilitating normal child development: thus, the unbiased germline mutation rate is unknown. To overcome this survivorship bias, one must examine a bulk sperm population comprised of reproductively “fit” and “unfit” gametes which accumulate DNMs amidst continuous cell division of spermatogonial stem cells. We hypothesized that the mutation rate in bulk sperm would be greater than pedigree-derived estimates, reflecting the continuous division of spermatogonial stem cells in a man’s lifetime. To test this, we used targeted Duplex DNA Sequencing to analyze low-frequency mutations in bulk sperm across 93 genes involved in spermatogenesis and DNA repair. This technology distinguishes true mutations as those present on complementary DNA strands from errors that arise on single strands. The resulting per-base error rate falls from 1/1000 to 1/100,000,000. In an ongoing study, we are applied Duplex Sequencing to cross-sectional and longitudinal bulk sperm samples from normozoospermic men (sperm concentration ≥30M/mL of ejaculate) aged between 24 to 68 years.
Our sequencing of bulk sperm supports the hypothesis that pedigree estimates of male germline mutation rates suffer from a survivorship bias. We observe an order of magnitude higher mutation rate (1.2×10-7) and an increase of 8 mutations/year in bulk sperm. Further, we find that an average of 6.25% of mutations are clonal (i.e., present in more than one SSC), and 7% of clonal mutations are pathogenic. Such clonal mutations confer a lifelong risk for the transmission of deleterious mutations.

Smart Cancer Genomics Technologies
Cancer is notoriously difficult to eradicate because tumors vary in their genetic makeup and in their susceptibility to cancer treatments. Furthermore, tumors adapt over time, allowing them to escape chemotherapies, metastasize, and ultimately end the patient’s life.
We have developed computational methods to track genetic changes in metastatic breast cancer tumors over the course of disease and treatment. Our goal is to help determine which genetic signatures predict aggressive growth or metastasis, which tumors respond to different treatments, and which changes signal emerging drug resistance. By following the evolutionary trajectories of individual tumors, we hope to develop precise and dynamic treatment regimens that will reduce unnecessary treatments and prevent disease relapse.
Browse our cancer genomics publications here.

Single Cell Sequencing Technology
Our goal is to identify targetable vulnerabilities for personalized medicine. We use state-of-the-art informatics tools to characterize patient tumors by analyzing multi-omics data including whole genome sequencing, bulk RNA sequencing and single cell RNA sequencing (scRNAseq) data. Specifically, scRNAseq, as a key part of our omics characterization platform, allows us to identify transcriptomic vulnerabilities from the pure tumor population without signal interference from other normal cells. From the omics data, we can identify the driver mutations and dysregulated pathways, and predict which drug(s) might work targeting these vulnerabilities. Then, integrating in vitro drug screen on patient tumor derived models, we are able to recommend drugs to oncologists. We have applied our precision oncology approach on advanced breast cancer patients and pediatric brain cancer patients in collaboration with Dr. Alana Welm, Dr. Sam Cheshier, and Dr. Philip Moos.
It’s known that within the tumor cell population, there might be different groups of cells that have different mutations or different gene expression. These groups of tumor cells are defined as subclones and cause the heterogeneity of the tumor. These subclone might have divergent behaviors during the tumor evolution and disease progression and have very different response to treatments. ScRNAseq allows us to dissect and study these subclones in isolation and provides unprecedented resolution to tumor heterogeneity. ScBayes, developed by Dr. Qiao in the lab, is a tool to integrate bulk DNA sequencing and scRNAseq data. Briefly, we can assign the subclone identity to single cells based on whether a cell contains variants specific to one subclone. Using this method, we studied the transcriptomic behavior of different subclones and their roles in tumor evolution along the treatment in chronic lymphocytic leukemia.
Infectious Disease and Metagenomics
Microorganisms inhabiting the human body outnumber human cells ten to one and are critical for our health and survival. Infectious disease is a global health problem, killing nearly 5 million children worldwide each year. Yet only a fraction of the microbes that promote human health or cause disease can be identified using traditional techniques.
In collaboration with ARUP Laboratories and IDbyDNA, Inc., we have developed Taxonomer, an ultrafast metagenomics tool that identifies all organisms present in a sample by rapidly analyzing the DNA and RNA sequences present. Taxonomer can identify microbes from many sample types, including body fluid, soil, food, and water. Taxonomer is already being deployed in University of Utah emergency room to diagnose respiratory infections, and for research analyses for children with Acute Respiratory Distress. Unlike other command-line metagenomics tools, Taxonomer is available over the web via iobio, and displays results in real time, using elegant graphics and interactive displays.
Browse our metagenomics publications here.
Learn more and try Taxonomer here.
Data Visualization
Computational genomics technologies have exploded in recent years, but most tools still require a bioinformatician and significant computational horsepower to use. We aim to change that with iobio, a web-based genomics tool suite that puts the power of a supercomputer into the hands of ordinary users.
The iobio platform allows researchers, clinicians, and data analysts to drive powerful genomic analyses themselves, using only a laptop computer. iobio analyzes genomic sequences in real time and generates interactive, visual, and intuitive displays. To date, four genomics applications have been deployed within iobio, including the metagenomics app Taxonomer, a variant prioritization app called gene.iobio, and two data quality inspection apps vcf.iobio and bam.iobio.
Browse our data visualization publications here.
Learn more and try iobio here.
Understanding Genomes

Genome Structure and Variation
Despite decades of effort, the genetic basis of most human diseases is poorly understood. Less than 5% of the heritability of most diseases can be explained by known genetic variation.
We develop tools and methods to discover, catalogue, and characterize genomic variation, including single nucleotide polymorphisms and structural rearrangements such as deletions, duplications, copy number variation, insertions, inversions and translocations. We also study the role of genomic variation in human health and disease.
Browse our genomic structure and variation publications here.

Genome Annotation
Accurate genome annotations form the basis for all future genetic and molecular biology research. The sequence of the first human genome took a decade and billions of dollars to assemble. Today, an individual lab can use our MAKER tool to assemble and annotate a new genome in a matter of hours, and the software is free for academic use.
MAKER and MAKER-P (optimized for plant genomes) have become the gold standard software for annotation of novel genomes, with hundreds of licenses issued worldwide. The tools have been used to annotate the genomes of the loblolly pine, the king cobra, the long-living sacred lotus, the spotted gar, the Burmese python, the desert woodrat, and dozens of others, nucleating entirely new research programs in the process.
Browse our genome annotation publications here.

Evolution of Genomes and Populations
Every genome is a treasure trove of information about the genetic and selective forces that drive molecular evolution and variation across and within species.
Our comparative genomics tools are used to identify molecular mechanisms of gene regulation and protein function, reveal ancient host-microbe conflicts, derive insights into global migration and population stratification, understand how mobile elements and viral infections generate genomic variation, and identify molecular changes underlying evolutionary transitions.
Browse our evolutionary genomics publications here.